I will perform scrnaseq data analysis
Postdoctoral Researcher in Bioinformatics
About this Gig
I will perform high-quality single-cell RNA-seq (scRNA-seq) data analysis using R packages such as Seurat, SingleR, SingleCellSignalR, and AUCell helping you uncover meaningful insights from your dataset.
Whether you are a student, researcher, or part of a biotech lab, I provide a reliable, reproducible, and publication-ready analysis workflow along with results interpretation.
My analysis includes:
- Quality control (QC) and filtering of cells and genes
- Data normalization, scaling, and integration (if needed)
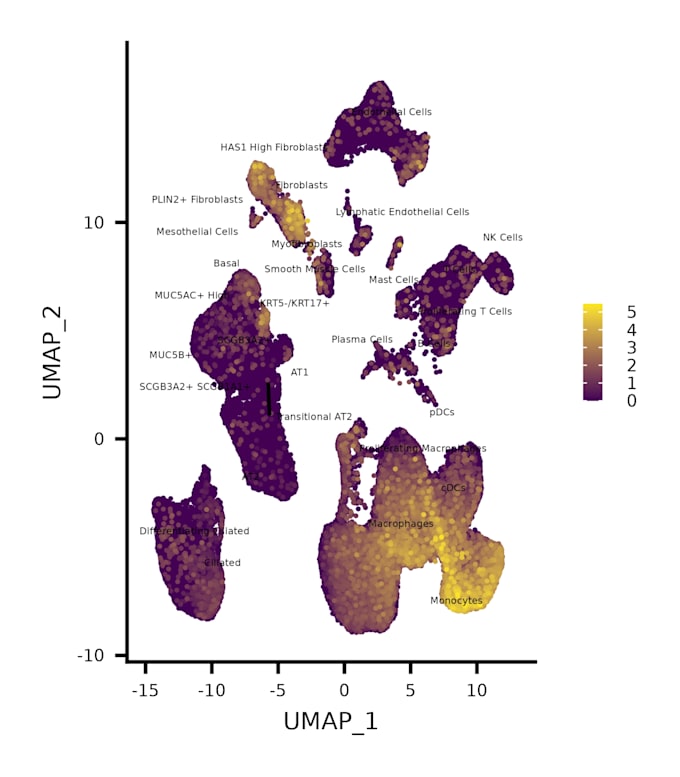
- Dimensionality reduction (PCA, UMAP/t-SNE)
- Cell clustering and marker gene identification
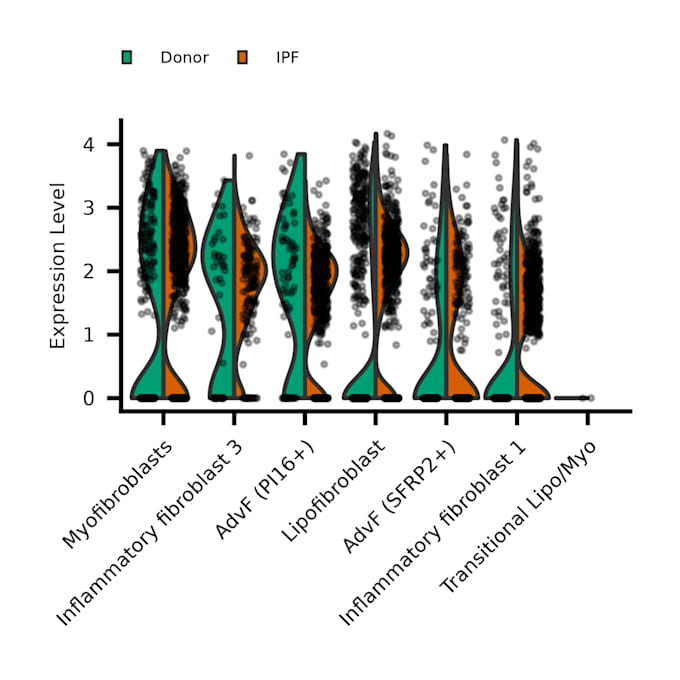
- Differential expression analysis between clusters or conditions
- Functional enrichment (GO/KEGG) of marker genes
- Publication-ready plots (UMAP/t-SNE, heatmaps, feature plots)
- Clear summary report and reproducible Seurat scripts.
I can work with raw count matrices or processed data.
If you have specific requirements, experimental conditions, or preferred reference genome, please contact me before ordering.
Deliverables are tailored to your dataset size and research goals, ensuring that your scRNA-seq results are ready for presentations, thesis, or publication.