
Mouna BS
PhD Drug Discovery Expert
Skills

See my services
Portfolio
Work experience

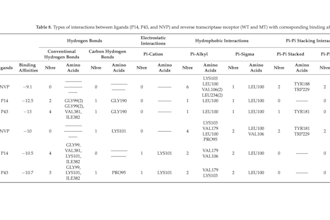
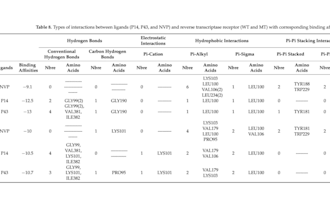
Published peer-reviewed research on HIV-1 drug discovery using advanced in silico techniques
Freelancer.com • Self-employed
Oct 2020 - Present • 5 yrs 7 mos
Work Experience Computational Chemist & Drug Design Researcher - Published peer-reviewed research on HIV-1 drug discovery using advanced in silico techniques. - Developed and validated QSAR (2D & 3D) models with strong predictive performance (R² up to 0.97). - Designed and optimized novel drug candidates targeting HIV-1 protease and reverse transcriptase. - Conducted molecular docking studies to evaluate ligand–protein binding affinities and interaction mechanisms - Performed molecular dynamics simulations (100 ns) to assess the structural stability of protein–ligand complexes - Applied ADMET and drug-likeness analysis to filter compounds with optimal pharmacokinetic profiles. - Successfully proposed new lead compounds with comparable or superior activity to existing drugs (e.g., darunavir benchmarks). - Used tools: Discovery Studio, MATLAB, XLSTAT, Gaussian, Avogadro, OpenBabel, ChemDraw ... Scientific Researcher – HIV-1 Inhibitor Design (Published Work) Academic Research Projects - Built robust QSAR models linking molecular descriptors to biological activity using PCA, MLR, and clustering methods. - Designed 35+ novel NNRTI compounds based on 3D-QSAR contour map insights. - Identified high-potential candidates with enhanced binding affinity and favorable ADMET properties. - Conducted statistical validation (LOOCV, external validation, Y-randomization), ensuring model reliability. - Evaluated drug resistance challenges and proposed solutions via structure-based design.